In the McLaughlin Lab we are committed to creating and maintaining a diverse, inclusive, and accessible training environment within our laboratory, department, and the Vassar College Community.
Additionally in Fall 2018, Prof. McLaughlin started an undergraduate STEM student group called the Alliance for Diversity Science and Engineering (ADSE) to support students here at Vassar, particularly students who are underrepresented in the STEM fields. Our undergraduate ADSE chapter is part of a national non-profit student run organization dedicated to building an inclusive and diverse community. Most ADSE chapters are for graduate students, however Vassar is home to one of only two undergraduate ADSE chapters in the country.
If you are interested in learning more about ADSE or about the projects/lab (after reading the research summaries below!), contact Dr. McLaughlin at kmclaughlin@vassar.edu.
Lab Publications (full list): NCBI Bibliography
Research Overview
We often think about bacteria only in the context of an infection, however the majority of bacteria we interact with are harmless- with many living symbiotically in/on us as part of the human microbiome. In the human gut alone there are more that 1200 different species of bacteria which help us stay healthy. When we are exposed to a bacterial pathogen that causes an infection, we take antibiotics which will kill the pathogen but unfortunately also eradicate the non-harmful bacteria.
How can we specifically target just the bacterial pathogen while leaving the symbiotes of our microbiome intact? Unfortunately, we do not currently have the technology or drugs available that can do this. Research in the McLaughlin lab aims to bridge the gap in knowledge that will allow us to one day make this a reality.
Our research is centered around the idea that a richer understanding of bacteria as a whole can help us to better target specific strains. We study uncharacterized proteins involved in various processes that are of importance to bacterial systems including antibiotic resistance transfer, carbohydrate utilization, and biosynthetic pathways like fatty acid synthesis.
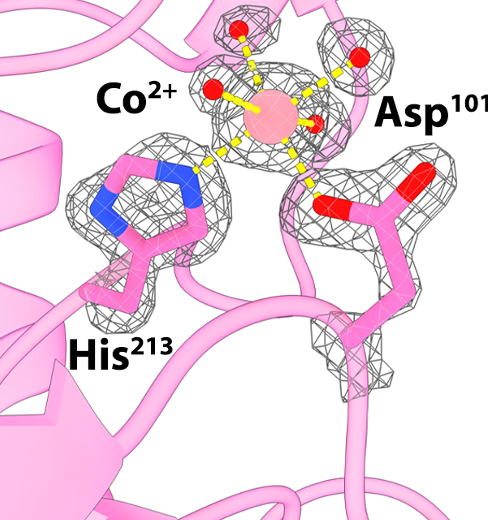
We use macromolecular x-ray crystallography to investigate protein structure and function. Coupled with molecular biology, biochemical, and biophysical techniques, it allows us to investigate structure-function relationship in various systems.
An overview of our general experimental strategy for studying target proteins is shown here:

Read more about each of the three research areas in the McLaughlin Lab here:
(1) spread of antibiotic resistance in Salmonella and Staphylococcus
(2) capsular polysaccharide biosyntheic enzymes from Bacteroides ovatus
(3) emerging pathogens proteome characterization






Bonus- Check out this video with Prof. McLaughlin discussing an introduction to protein structure and a peek into the research in the McLaughlin Lab: https://vimeo.com/450228303/281c87d3ac